Advancing Cellular Imaging with Digital, User-Friendly Systems
Theodore Price, PhD, talks to contributing editor Tanuja Koppal, PhD, about the advent of digital cell imaging instruments that are very mobile, user-friendly, inexpensive, and intuitive to work with.

Theodore Price, PhD, associate professor in the School of Behavioral and Brain Sciences at University of Texas at Dallas, talks to contributing editor Tanuja Koppal, PhD, about the advent of digital cell imaging instruments that are very mobile, user-friendly, inexpensive, and intuitive to work with. He recommends their use at undergraduate institutions where a lot of students need to be trained in a short period of time. These instruments are also good as screening tools or for proof-of-concept studies to show that an experiment works before it is performed on high-cost, high-powered confocal imaging systems.
Q: Can you give us some details about your research and how you use cellular imaging?
A: We are a neuroscience and pharmacology laboratory, and our area of focus is pain and finding out how pain becomes chronic. We try to find targets that we think can manipulate pain plasticity so we can develop therapeutics that can permanently reverse chronic pain states. In order to accomplish that, we do behavioral animal studies and a variety of cellular signaling assays. We do a lot of imaging in primary cell cultures, namely dorsal root ganglion neurons derived from either wild-type mice or a variety of transgenic mice. We are interested in looking not only at cellular signaling events, but also events taking place in subcellular compartments. One of the main events that we study is translational control or protein synthesis. In the last couple of years, the number of methods that are available to study translational control in subcellular compartments has increased. We do a fair amount of confocal imaging using antibodies or cellular reporters. We have also started to do some live cell imaging.
Our typical workflow involves generating cell cultures from mice, then using pharmacologic or genetic approaches to manipulate the pathways that we are interested in, and then imaging those pathways. We previously used confocal imaging to do everything—from looking at cell staining to getting the images for publication purposes—but that was a slow process. Now we use a digital, handheld fluorescent cell imager to look at cells before we use the confocal system. Once we know that our experiment has worked—the cells look healthy, the antibody has stained well, and other things that we need to check look good on the digital cell imager—we move on to the more expensive confocal machine.
Q: What were your biggest challenges doing cell imaging, and how has the process changed?
A: Until three or four years ago, a lot of our time was spent using Western blotting (WB) to look at phosphoproteins. WB is easy, but it’s slow and it does not give you any type of cellular resolution. A few factors have facilitated the change in how we do things. We now have a broad range of phospho-specific antibodies that work very well for immunohistochemistry or immunocytochemistry. We try to take advantage of transgenic reporters that are available. These are mice expressing green fluorescent protein that are then tagged to certain proteins involved in translational control, so we can see where these proteins are going in the cell. We also use a number of new assays such as Click Chemistry and the SUnSET technique to measure nascent protein synthesis. With these new assays and imaging techniques, we can see not only what types of changes are occurring, but also where in the cell they are happening.
Q: What are the advantages of using a digital fluorescent cell imager?
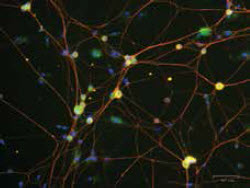 Mouse dorsal root ganglion neurons were cultured for five days and stained for rck (green) labeling p bodies, beta III-tubulin (red) labeling neurons, and dapi (blue) labeling nuclei.Image courtesy of Jamie Moy, taken on ZOE™ Fluorescent Cell ImagerA: We are an undergraduate institution, so besides having graduate students and postdoctoral fellows, we have a lot of undergraduate students working in our labs. These students have no experience doing cell imaging, and although they can be trained, it’s a little nerve-racking to have them start working on a $300,000 confocal microscope. The digital cell imager is a small epifluorescent microscope that is relatively new and inexpensive. Using this, the students can hone their skills and see whether their experiment is working, and it provides very good images given its size and cost. It’s essentially unbreakable as we move it around the lab from one desk to another. We typically use this cell imager to see if we have good quality of cells, if the treatment worked, or if the antibody staining is good before we start working on the confocal microscope. In some cases, we also change our workflow so we can do all imaging experiments on the digital imager and not have to use the confocal microscope. Its sensitivity is good enough to do some simple assays, and it has freed up the time on our confocal microscope, which was really custom-built to do live cell imaging. Having both the confocal microscope and the epifluorescence imager available helps students understand what each instrument is really meant for, and makes it possible for us to utilize them both to the best of their abilities. You can get beautiful images in a couple of hours using the confocal microscope, but you can get the same answer in just a few minutes using digital imaging. You can then move on to your next experiment having saved both time and money. By pushing the envelope on the specificity of the reporters we make, we are able to spend more time working on the technology and working on new drug targets.
Mouse dorsal root ganglion neurons were cultured for five days and stained for rck (green) labeling p bodies, beta III-tubulin (red) labeling neurons, and dapi (blue) labeling nuclei.Image courtesy of Jamie Moy, taken on ZOE™ Fluorescent Cell ImagerA: We are an undergraduate institution, so besides having graduate students and postdoctoral fellows, we have a lot of undergraduate students working in our labs. These students have no experience doing cell imaging, and although they can be trained, it’s a little nerve-racking to have them start working on a $300,000 confocal microscope. The digital cell imager is a small epifluorescent microscope that is relatively new and inexpensive. Using this, the students can hone their skills and see whether their experiment is working, and it provides very good images given its size and cost. It’s essentially unbreakable as we move it around the lab from one desk to another. We typically use this cell imager to see if we have good quality of cells, if the treatment worked, or if the antibody staining is good before we start working on the confocal microscope. In some cases, we also change our workflow so we can do all imaging experiments on the digital imager and not have to use the confocal microscope. Its sensitivity is good enough to do some simple assays, and it has freed up the time on our confocal microscope, which was really custom-built to do live cell imaging. Having both the confocal microscope and the epifluorescence imager available helps students understand what each instrument is really meant for, and makes it possible for us to utilize them both to the best of their abilities. You can get beautiful images in a couple of hours using the confocal microscope, but you can get the same answer in just a few minutes using digital imaging. You can then move on to your next experiment having saved both time and money. By pushing the envelope on the specificity of the reporters we make, we are able to spend more time working on the technology and working on new drug targets.
Q: What were you looking for when you invested in this new system?
A: We were looking to get an epifluorescent microscope that we could use at the end of our experiment but before we started our work on the confocal instrument. I did see another system a couple of years ago, but it did not have all the intangibles that we were looking for, so it was not worth the investment. With the recent instrument, I was struck by how mobile these imagers had become. It really did not need any dedicated bench space and the images we obtained were instantaneous and quite beautiful. Using it is totally intuitive, with a touch screen interface and the ease of a personal tablet—any 18- to 25-year-old student can be taught how to use it in a few minutes. That’s very valuable to us since we have a lot of undergraduates in our lab who need to be taught things very quickly.
Q: Can you elaborate on the intangibles that you were looking for?
A: The key factor for us was mobility. We needed an instrument that we could move around the lab and one that had lots of different uses. The other factor was ease of use. This digital imaging system is really intuitive, and you can figure out how to use it almost instantly. If you have a smartphone, you will know how to use it.
Q: What kind of improvements are you looking for in these new imaging systems?
A: The idea of using very mobile, all-digital microscopes is just catching on, and there is definitely a lot of utility for them. There is a good opportunity to expand the optics on this instrument to give it more flexibility. I am sure it’s possible to add to the magnification and other optical features without changing the mobility and ruggedness of the instrument. Even if these improvements add ~$5,000 more to the cost, it still remains pretty affordable. Also, adding more flexibility in terms of the epifluorescence will add more opportunity in terms of the applications. Initially we were not quite sure how this instrument would be used in our lab, but now we find it being used all the time. People use it just to look at their cells and make sure they look healthy. I can see us having one of these digital systems in the tissue culture room because it’s cheaper than the inverted microscope we have that nobody uses anymore.
Q: With this instrument, are you compromising on the throughput of your assays?
A: The throughput of the digital imaging system is very good for what we use it for, which is looking at cells on coverslips. We have tried to use it for 96-well plate imaging but we can’t quite image an entire well at a time. The ability to image a 96-well plate would definitely increase its appeal. We could then do high-throughput imaging assays in a short amount of time.
Q: Do you have any advice for lab managers looking to evaluate these digital microscopes?
A: If you do a lot of confocal imaging, then this system acts as a good screening tool. If you are in an undergraduate institution or a medical school, where you need to train students quickly and make them more productive, then a digital system can be a good way to get started. Getting people trained and getting productivity up is important to all labs.
Dr. Theodore (Ted) Price is an associate professor in the School of Behavioral and Brain Sciences at University of Texas at Dallas. He received his PhD in 2003 from the University of Texas Health Science Center at San Antonio under the mentorship of Christopher Flores and Kenneth Hargreaves, and was a postdoctoral fellow in the laboratory of Fernando Cervero at McGill University until 2007, where he developed the first studies on how local control of protein synthesis in neurons is involved in the development of chronic pain disorders. He was a faculty member of The University of Arizona School of Medicine from 2007 to 2014, where he further developed work on nervous system plasticity in relation to pain with funding from the National Institutes of Health, the Rita Allen Foundation, and the Migraine Research Foundation. Dr. Price is the recipient of Young Investigator awards from the American Pain Society and International Association for the Study of Pain, and has published more than 70 peer-reviewed articles in international journals.
