Missouri S&T Chemist Rolls the Dice to Better Identify Chiral Molecules in Drugs
“High risk, high reward” is the kind of discovery Dr. Garry Grubbs seeks with a new experiment designed to rapidly identify the atomic structure of chiral molecules widely used in pharmaceutical drugs
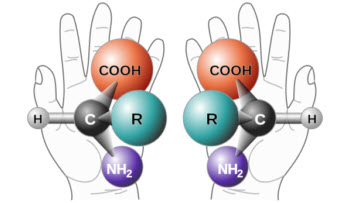 The atomic structure of chiral molecules resembles "handiness."Credit: Wikimedia COmmons
The atomic structure of chiral molecules resembles "handiness."Credit: Wikimedia COmmons
“High risk, high reward” is the kind of discovery Dr. Garry Grubbs seeks with a new experiment designed to rapidly identify the atomic structure of chiral molecules widely used in pharmaceutical drugs. The finding could significantly reduce the time and costs involved in pharmaceutical development and manufacturing.
Grubbs, an assistant professor of chemistry at Missouri S&T, received nearly $60,000 in July from the National Science Foundation as an early-concept “EAGER” grant for his experiment. EAGER projects explore untested, but potentially transformative research ideas considered “high risk-high payoff” because they involve radically different approaches or engage novel perspectives.
Chiral molecules possess identical atomic connectivity, but their atoms are arranged in mirror-image configurations that cannot be superimposed on each other. Often described as “handedness,” the atomic structures of chiral molecules contain similar physical properties (as do the mirror images of the right and left hand), but their chemical properties usually differ.
Even though chiral molecules are nearly physically identical, life cells chemically respond with the desired pharmacological effect to only one of the mirrored molecular structures, making their identification a crucial task.
By some estimates, 90 percent of today’s drugs are chiral. And at present, those molecules are cumbersome to identify.
“The pharmaceutical industry doesn’t have an efficient technique to discern the chirality of certain molecular structures used in drug development,” says Grubbs. “Because a lack of discernment can lead to dangerous physiological effects in medicines, much time and care is now being taken to ensure the correct chiral form is used in drug synthesis.”
Grubbs says the pharmaceutical industry spends billions of dollars a year in identifying chirality through several time-consuming methods in order to avoid synthesizing the wrong molecular structure.
Current techniques include traditional and vibrational circular dichroism, optical rotation, nuclear magnetic resonance and x-ray crystallography—then separating the desired chiral structure from the undesired one by gas or liquid chromatography. Grubbs says the combination of these techniques can take many hours where a standardized analysis used in their place could take under a minute.
Grubbs has proposed a method to speed up this process by using an existing, advanced microwave spectroscopy technique known to distinguish molecules in three-dimensional space on molecular effects generated by singular atomic nuclei. Based on these interactions, he seeks to tease out and gather the information to identify chiral molecules.
Because these observations can occur in less than a minute, while also quantifying the reaction without the need to separate the reaction mixture, Grubbs believes it will drastically speed up chiral discernment.
“Monitoring molecular geometry based on singular atomic effects is a new, innovative approach, and coupling the effect of atomic interactions with chirality is a niche area,” says Grubbs. “If we can distinguish chirality in a singular process, it can potentially be automated, and the implications would expand into new areas.”
“Along with others in this area, I’m working to totally change our way of thinking when it comes to experimentally identifying chirality,” says Grubbs.
View the abstract: Three-wave mixing techniques to study and utilize nuclear quadrupole coupling effects in chiral molecules.
