Shape and Size of DNA Lesions Caused by Toxic Agents Affects Repair of DNA
SDSC’s ‘Gordon’ helps describe complex process that’s fundamental to cancer initiation
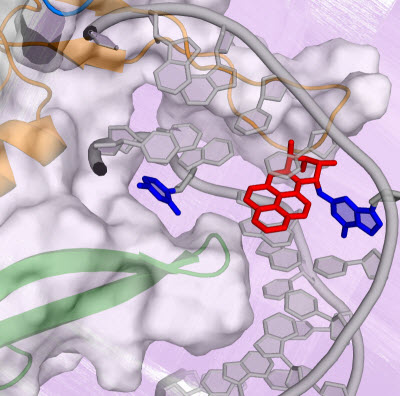 Conformational capture of a pre-flipped base by the NER lesion recognition protein yeast-XPC/Rad4.Graphic courtesy of Hong Mu, New York University.Every day our bodies come under a barrage of toxic agents—cigarette smoke, the sun, free radicals, and other carcinogenic substances—that create damaging lesions in our DNA that can initiate cancer and other human diseases.
Conformational capture of a pre-flipped base by the NER lesion recognition protein yeast-XPC/Rad4.Graphic courtesy of Hong Mu, New York University.Every day our bodies come under a barrage of toxic agents—cigarette smoke, the sun, free radicals, and other carcinogenic substances—that create damaging lesions in our DNA that can initiate cancer and other human diseases.
Fortunately, nature has provided living organisms with repair processes to seek out and remove such dangerous lesions; repair allows the DNA to be restored to its original base sequence so it can carry out its fundamental jobs: to be replicated and to be copied into a sister molecule, RNA, for the manufacture of proteins and other tasks.
The 2015 Nobel Prize in Chemistry was awarded to three scientists for their work on the mechanisms of DNA repair. Nonetheless, much remains to be understood about these intricate molecular processes.
Now, a team led by professor Suse Broyde and postdoctoral researcher Hong Mu—both in the Department of Biology at New York University (NYU)—has identified and described how a major player in the repair process, called nucleotide excision repair or NER, works to recognize certain lesions for subsequent removal by the NER machinery. Included among the lesions are bulky chemicals that link tightly to the DNA bases; these are called polycyclic aromatic hydrocarbons (PAH), or carcinogenic precursors inhaled through automobile exhaust or cigarette smoke, that ultimately can attach to our DNA.
The findings, published as an “Editor’s Choice” article in a recent issue of Chemical Research in Toxicology, a journal of the American Chemical Society, are helping researchers to better understand why certain lesions triggered by environmental and other agents are more likely to be repaired while others persist to cause mutations and cancer.
 DNA Basics: DNA is comprised of two inter-wound strands that form a double helix. The rungs of the double helical ladder contain four chemical moieties called bases, namely A (adenine), G (guanine), C (cytosine), and T (thymine), and in the two strands each A is paired by a T and each G is paired by a C; these are known as complementary base pairs. When the DNA is copied during replication, the two strands unwind and each daughter strand is the complement of the mother strand. When this process is inaccurate through damage by toxic agents, mutations result, which can initiate cancer and other human diseases.Image courtesy of the San Diego Supercomputer Center
DNA Basics: DNA is comprised of two inter-wound strands that form a double helix. The rungs of the double helical ladder contain four chemical moieties called bases, namely A (adenine), G (guanine), C (cytosine), and T (thymine), and in the two strands each A is paired by a T and each G is paired by a C; these are known as complementary base pairs. When the DNA is copied during replication, the two strands unwind and each daughter strand is the complement of the mother strand. When this process is inaccurate through damage by toxic agents, mutations result, which can initiate cancer and other human diseases.Image courtesy of the San Diego Supercomputer Center
DNA repair begins with a protein called XPC (xeroderma pigmentosum C protein complex), whose job is to patrol the genome for certain types of lesion-induced DNA disturbances. Once it encounters a damaged DNA, it inserts a beta-hairpin (a simple protein structural motif involving two beta strands that look like a hairpin) between the two DNA strands, which serves to recognize the lesion so that NER can ultimately remove it.
“It has been shown that NER recognizes a wide range of DNA lesions, whose NER susceptibilities vary for reasons that are not well understood,” said Broyde. “Understanding differences in the XPC recognition mechanism for lesions with different sizes and shapes may help us understand this variability.”
In the current study, the researchers sought to identify and describe the molecular pathway that Rad4, a yeast version of XPC, takes when it binds to a bulky DNA lesion derived from the PAH benzo[a]pyrene. For this DNA lesion, a large multi-ringed structure is bound to the DNA base guanine (G); the rings are inserted into the DNA helix, rupturing the DNA base pairs. The partner base cytosine (C) is extruded from the double helix.
Atomic-level Simulations Using Gordon
To learn more about the binding process of the yeast-XPC, called Rad4, the researchers turned to the Gordon supercomputer at the San Diego Supercomputer Center (SDSC) at UC San Diego to simulate at an atomic level the lesion recognition pathway. As described in the paper, the simulations showed a pathway in which the yeast-XPC/Rad4 initially captures the extruded/pre-flipped base C, the base partner to the damaged G.
Subsequently, as the DNA bends and unwinds, a second base is flipped into the protein, the beta-hairpin is inserted into the double helix, while the multi-ringed part of the lesion is pushed to the helix exterior. The lesion is eventually excised by the NER machinery.
The mechanism differed significantly from that of a prior study with a UV-light-induced lesion, CPD, which can cause skin cancer. This lesion-containing DNA duplex does not contain an extruded/pre-flipped base that can be captured initially. In this case, the simulations showed that two bases opposite the lesion flipped in correlated motion to open the lesion site for subsequent beta-hairpin insertion, while the small CPD lesion is easily extruded before the hairpin inserted.
“Combined, our research shows that the structure of the lesion-containing DNA impacts the binding pathway of yeast-XPC/Rad4 to the lesions,” said Mu, the study’s first author, “and that the pathway may be tailored to the particular disturbance to DNA caused by lesions of various sizes and shapes. This capability may play an important role in the ability of XPC to recognize a wide variety of lesion types. In the case of repair resistant lesions it is hypothesized that productive binding that leads to subsequent excision is obstructed.”
Added Broyde: “Individuals who harbor repair-resistant lesions could potentially be identified through highly sensitive measuring techniques (“adductomics”) that can utilize, for example, hair or urine samples. Such individuals could then be counseled to alter their lifestyles, to cease smoking for example, and be vigilant about monitoring early cancers.”
The work also has application to drug design. In the case of chemotherapeutic drugs, such as cis-platinum, the efficacy of the drug is diminished by its repair through NER; a goal in the design of more advanced agents of this family is to develop ones that are less repair-susceptible. Understanding the essential mechanism that initiates NER through lesion recognition could aid in the design of more potent drugs that are NER resistant.
The simulations were based on molecular dynamics calculations designed to find the most efficient pathway used by XPC/Rad4 to move from a state where its lesion-recognition domain was unbound to where it is correctly bound to the DNA lesion. Since the pathway was totally unknown, the researchers said it was essential to explore many possibilities to locate the best path, with the lowest and hence most favorable energy barrier for binding.
“These extremely compute-intensive calculations required the resources of Gordon in order to permit the computations to be carried out in parallel and hence be achieved in a reasonable amount of time,” said Mu.
As a next step, the researchers are preparing to investigate a library of DNA lesions characterized by Nicholas Geacintov, professor of chemistry at NYU and co-author of the paper. Geacintov, in collaboration with professor Dinshaw Patel of Memorial Sloan Kettering Cancer Center, and Broyde, has determined the NMR structures, NER efficiencies, and other biochemical and biophysical properties of the damaged DNA duplexes in the library. This work will seek to evaluate how pathways differ among a variety of lesions and evaluate whether the pathways correlate with repair susceptibility.
Also participating in the study were Jung-Hyun Min, associate professor of chemistry, University of Illinois at Chicago; and Yingkai Zhang, professor of chemistry at NYU.
Support for the study came from the National Institute for Environmental Health Sciences (NIEHS), National Institute of General Medical Sciences (NIGMS), and the National Science Foundation (NSF), grant # MCB060037 for the computing resources at XSEDE.
