Researchers at the U.S. Department of Energy’s (DOE's) Argonne National Laboratory have mapped out two very different types of protein. One helps soil bacteria digest carbon compounds; the other protects cells from the effects of harmful molecules.
In order to uncover the structure of these proteins, researchers used a technique called protein crystallography. Like a mosquito trapped in amber, compounds that are crystallized are placed in array in identical positions and ordered so that scientists can target them with X-ray beams and work backwards from the scattering patterns produced to recreate their three-dimensional structures atom by atom.
Bacterial breakdown
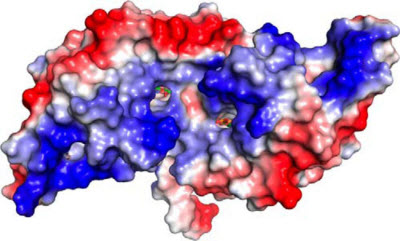 Researchers at Argonne modeled the HcaR protein complex, above, a sort of molecular policeman that controls when to activate genes that code for enzymes used by Acinetobacter bacteria to break down compounds for food. Understanding these processes can help scientists develop ideas for converting more carbon in soil.Image courtesy Kim et al./Journal of Biological ChemistryIn the first study, a group of researchers from the Structural Biology Center, which is funded by DOE’s Office of Science, mapped out a protein responsible for breaking down organic compounds in soil bacteria, an important process for recycling carbon in the ecosystem.
Researchers at Argonne modeled the HcaR protein complex, above, a sort of molecular policeman that controls when to activate genes that code for enzymes used by Acinetobacter bacteria to break down compounds for food. Understanding these processes can help scientists develop ideas for converting more carbon in soil.Image courtesy Kim et al./Journal of Biological ChemistryIn the first study, a group of researchers from the Structural Biology Center, which is funded by DOE’s Office of Science, mapped out a protein responsible for breaking down organic compounds in soil bacteria, an important process for recycling carbon in the ecosystem.
The bacteria used, called Acinetobacter, is located mostly in soil and water habitats, where it helps to change aromatic compounds (named for their ring shape) into forms that can be used as food.
One of the sources of aromatic compounds found in soil is lignin, a tough polymer that is an essential part of all plants and that’s hard for many organisms to digest.
“But Acinetobacter can utilize these aromatic compounds as their sole source of carbon,” said Andrzej Joachimiak, who co-authored both studies and is the director of the Structural Biology Center and the Midwest Center for Structural Genomics at Argonne.
In order for Acinetobacter to break down the aromatic compounds, it needs to produce catabolic enzymes, molecular machines built from an organism’s DNA that break down molecules into smaller parts that can be digested.
Whether or not membrane transporters and catabolic enzymes are produced falls to the HcaR regulator, a sort of molecular policeman that controls when the genes that code for these enzymes can be activated.
Joachimiak and his colleagues found that the regulator works in a cycle, activating genes when aromatic compounds are present and shutting genes down when the compounds are used up.
“By nature it is very efficient,” Joachimiak said. “If you don’t have aromatic compounds inside a cell, the operon is shut down.”
The research team didn’t stop at mapping out the regulator itself; to discover how the cycle worked, they crystalized the HcaR regulator during interactions with its two major inputs: the aromatic compounds and DNA.
The group found that when aromatic compounds are not present in the cell, two wings found on either side of the HcaR regulator wrap around the DNA. This action is mirrored on both sides of the regulator, covering the DNA regulatory site and preventing genes from being activated.
“This is something that has never been seen before,” Joachimiak said.
When the aromatic compounds are present, however, they attach themselves to the HcaR regulator, making it so stiff that it can no longer grapple with the DNA.
Joachimiak said that this knowledge could help outside of the lab, with applications such as a sensor for harmful pesticides and as a template for converting more carbon in soil.
“If we can train bacteria to better degrade lignin and other polymers produced by plants during photosynthesis, more natural carbon sources can be utilized for example for production of biofuels and bioproducts,” Joachimiak said.
The paper was published earlier this year by the Journal of Biological Chemistry under the title “How Aromatic Compounds Block DNA Binding of HcaR Catabolite Regulator.” It was supported by the National Institutes of Health and the U.S. Department of Energy (Office of Biological and Environmental Research).
Protective proteins
A second paper focuses on a family of proteins identified as DUF89, which stands for “domain unknown function.” This family is conserved across all three branches of the phylogenetic tree, which means that it is likely essential to many life forms.
DUF89 has been identified as a type of enzyme called a phosphatase, which strips molecules of their phosphate groups. The paper’s authors hypothesized that DUF89 proteins use this ability to save useful proteins in a cell from rogue molecules which could alter their structure, making them useless or destructive.
The study found that DUF89 proteins use a metal ion, probably manganese, to lure in potentially harmful molecules and a water molecule to break off their phosphate group.
DUF89 proteins could have an important role in breaking down a specific type of disruptive molecule: sugar. When the concentration of sugar in blood reaches high levels, simple sugars can have unwanted side reactions with proteins and DNA through a process called glycation.
“We always have to deal with these side reactions that happen in our cells, and when we get older, we have an accumulation of these errors in our cells,” Joachimiak said.
Joachimiak said that this research could help scientists develop DUF89 treatments from non-human sources as a way to combat glycation in the bloodstream.
The paper was published on the Nature Chemical Biology website on June 20 under the title “A family of metal-dependent phosphatases implicated in metabolite damage-control.” Other authors on the paper were from the University of Florida, the University of Toronto, the University of California-Davis and Brookhaven National Laboratory. It was supported by the National Science Foundation, Genome Canada, the Ontario Genomics Institution, the Ontario Research Fund, the Natural Sciences and Engineering Research Council of Canada, the National Institutes of Health, the C.V. Griffin Sr. Foundation, and the U.S. Department of Energy (Office of Basic Energy Sciences and Office of Biological and Environmental Research).
Both studies used X-rays from the Advanced Photon Source, a DOE Office of Science User Facility, using beamlines 19-ID and 19-BM.
Both also stem from the goal of the Midwest Center for Structural Genomics, which is to discover the structure and function of proteins potentially important to biomedicine.
Joachimiak said that despite the new findings from these studies, when it comes to understanding what proteins do, we still have a long way to go.
“When we sequence genomes, we can predict proteins, but when we predict those sequences we can only say something about function for about half of them,” Joachimiak said.
















