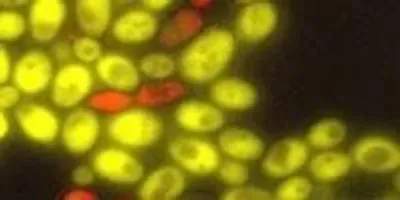
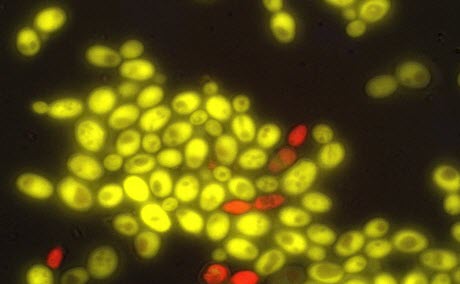
 UW researchers have produced cell-to-cell communication in baker’s yeast, a first step in building multicellular organisms from scratch. The red yeast cells produce a plant hormone, which “tells” the green cells to express a gene differently.Image credit: University of WashingtonNow, a team of University of Washington researchers has engineered yeast cells (Saccharomyces cerevisiae) that can “talk” to one another, using a versatile plant hormone called auxin. In a paper published June 23 in the American Chemical Society’s journal ACS Synthetic Biology, the researchers describe a novel cell-to-cell communication system that enables one yeast cell to regulate the expression of genes and influence the behavior of an entirely separate yeast cell.
UW researchers have produced cell-to-cell communication in baker’s yeast, a first step in building multicellular organisms from scratch. The red yeast cells produce a plant hormone, which “tells” the green cells to express a gene differently.Image credit: University of WashingtonNow, a team of University of Washington researchers has engineered yeast cells (Saccharomyces cerevisiae) that can “talk” to one another, using a versatile plant hormone called auxin. In a paper published June 23 in the American Chemical Society’s journal ACS Synthetic Biology, the researchers describe a novel cell-to-cell communication system that enables one yeast cell to regulate the expression of genes and influence the behavior of an entirely separate yeast cell.
It’s a basic step in understanding the communication and cooperative processes that might lead to synthetic stem cells that could grow into artificial organs or organisms that require different types of cells to work together.
“Until you can actually build a multicellular organism that starts from a single cell, you don’t really understand it. And until we can do that, it’s going to be hard to do things like regrow a kidney for someone who needs it,” said senior author Eric Klavins, a UW associate professor of electrical engineering and of bioengineering.
It might also enable engineered yeast to perform complicated behaviors that coordinated multicellular systems such as our immune system can pull off, like recognizing an invading pathogen and mounting a response. If so, one might program those cells to collaboratively diagnose the flu or malaria: just add saliva to a packet of yeast and see if it changes color.
For now, though, the team spearheaded by lead authors Arjun Khakhar, a UW doctoral student in bioengineering, and Nicholas J. Bolten, a UW doctoral student in electrical engineering, simply wanted to see if it could induce one yeast cell to send a signal that sets off a cascade of changes in another cell.
In the initial experiment, they used the plant hormone auxin — which yeast cells don’t normally recognize or respond to — to “turn off” a target gene in another cell. In this case, the gene that was switched off was an inserted jellyfish gene that turned the yeast fluorescent green.
“This project was to find out whether we could use auxin to make the cells talk to each other in a really simple way,” said Klavins. “We’re not sending complicated messages yet. One cell is saying ‘hello?’ and the other cell says ‘I can hear you.’ Eventually they’ll say ‘I’m this kind of cell. What are you? Let’s work together.’ But for now it’s pretty much ‘hi.'”
Synthetic biologists, who assemble genetic parts in new ways with the goal of popping them into an organism to produce reliable behaviors, have struggled to build modules that enable cell-to-cell communication in organisms that don’t naturally do it.
The UW team overcame this hurdle by engineering a suite of novel transcription factors — proteins that control whether a specific gene inside a cell’s DNA is expressed or not — with varying sensitivities to auxin. That “tunability” offers important control in regulating cell behavior.
With co-author and UW associate biology professor Jennifer Nemhauser, the UW team figured out how to make a “sender” yeast cell produce auxin, a versatile hormone that controls everything from where a plant’s roots develop to how effectively they fight off pathogens. Through trial and error, the team learned an enzyme borrowed from a soil bacterium can induce yeast to convert a commonly available chemical into auxin.
In the “receiver” yeast cells, the researchers inserted the new transcription factor — which was assembled from so many different genetic parts that they call it the “Frankenfactor” – and engineered it to activate the jellyfish gene that turned the cell green.
When the sender cell released auxin, additional proteins that the researchers introduced in the receiver cell were able to degrade the Frankenfactor and switch off the gene that turned the receiver cell green.
 On the left, a transcription factor designed by UW researchers (gray and red) is integrated into the “receiver” yeast cell. This activates a gene that turns the receiver cell fluorescent green. On the right, a “sender” yeast cell produces a plant hormone called auxin (orange dots). In the receiver cell, auxin recruits another protein (blue) to disable the transcription factor and turn off the gene that turned the receiver cell green.Image credit: University of WashingtonThat type of simple communication forms the bedrock of multicellular organisms in which different types of cells collaborate to carry out complicated tasks. As a next step, the UW team plans to test whether auxin can induce more complex behaviors in yeast cells, such as forming patterns or cooperatively computing basic functions.
On the left, a transcription factor designed by UW researchers (gray and red) is integrated into the “receiver” yeast cell. This activates a gene that turns the receiver cell fluorescent green. On the right, a “sender” yeast cell produces a plant hormone called auxin (orange dots). In the receiver cell, auxin recruits another protein (blue) to disable the transcription factor and turn off the gene that turned the receiver cell green.Image credit: University of WashingtonThat type of simple communication forms the bedrock of multicellular organisms in which different types of cells collaborate to carry out complicated tasks. As a next step, the UW team plans to test whether auxin can induce more complex behaviors in yeast cells, such as forming patterns or cooperatively computing basic functions.
Since auxin is a plant hormone, mammalian cells also ignore it, making auxin a potentially useful tool in designing gene therapies or other applications without adverse reactions in humans. The UW method, which uses a “guide RNA” to target the gene of interest, could be adapted to produce a number of genetic or behavior changes.
“If you ask someone in computer science what they can do with a programming language, they’ll laugh and say they can do anything with it,” Klavins said. “If we can figure out the programming language of life, we can do anything that life does — except in a more controllable, reliable way.”
The research was funded by the National Science Foundation and the Paul Allen Family Foundation.